- Blog
- Clockit easy schedule creator
- Cateye cafe
- Minotaur maze key grounded
- Yu zhang ivinci health
- Deathmetal reddit
- Forza motorsport 8 toyota coming back
- Mass watermark photoshop
- Atc 250r for sale near me
- Histera transracial
- A wimoweh meaning
- Positively charged amino acids
- Power tae fresh download
- Lightzone vignette
- Plants to cover wall
- Macbook air flash storage firmware update 1-1- download
While it is known that there is great variation in ribosomal velocity along even a single transcript, what determines how fast a transcript (or part thereof) is processed is unresolved. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.Ĭompeting interests: The authors have declared that no competing interests exist. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.įunding: LDH is a Wolfson Royal Society Research Merit Award Holder. Received: JAccepted: FebruPublished: March 12, 2013Ĭopyright: © 2013 Charneski, Hurst. PLoS Biol 11(3):įred Hutchinson Cancer Research Center, United States of America A missed stop codon or frameshifting would then lead to a stalled ribosome, which is consistent with experimental data.Ĭitation: Charneski CA, Hurst LD (2013) Positively Charged Residues Are the Major Determinants of Ribosomal Velocity. Finally, we observe that the poly-A tail provides a massively positively charged terminus no matter in which frame it is translated. We also show why, if protein charge were not considered, one could be misled into suspecting a role for non-optimal codons. We show that positive charge explains the sites where ribosomes stall most commonly within transcripts. We find on the contrary that it is not RNA-level features but positive charges in the already translated protein that most retard ribosomes, possibly by interacting with the negatively charged ribosomal exit tunnel.
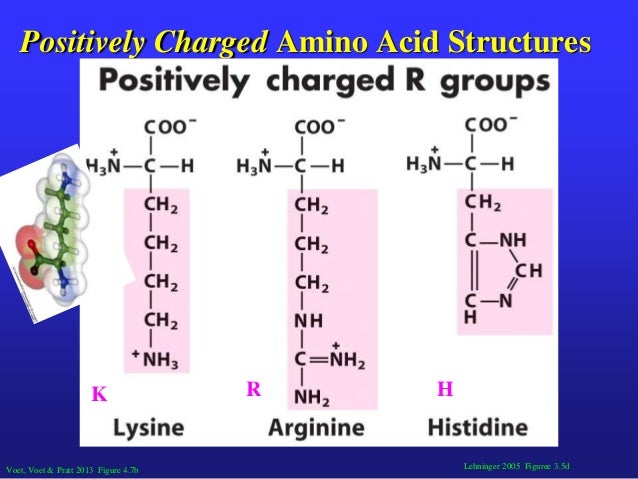
It has long been thought that RNA-level features modulate translation rates, whether by delays incurred through the presence of codons that require relatively rare tRNAs, or by regions of mRNA folding that physically impede ribosomal progression. Ribosomes do not synthesize protein at a constant rate along transcripts, and changes in translation speed can have knock-on consequences for the expression of that protein, even altering its folding or subcellular localization. We thus expect that a translated poly-A tail, encoding for positively charged lysines regardless of the reading frame, would act as a sandtrap for the ribosome, consistent with experimental data. The effect of charged amino acids is additive, with ribosomal occupancy well-predicted by a linear fit to the density of positively charged residues. Such slowing is independent of and greater than the average effect owing to mRNA folding. By contrast, we find that positively charged amino acids greatly retard ribosomes downstream from where they are encoded, consistent with the suggestion that positively charged residues interact with the negatively charged ribosomal exit tunnel.

While codon usage is classically considered a major determinant, we find no evidence for this.
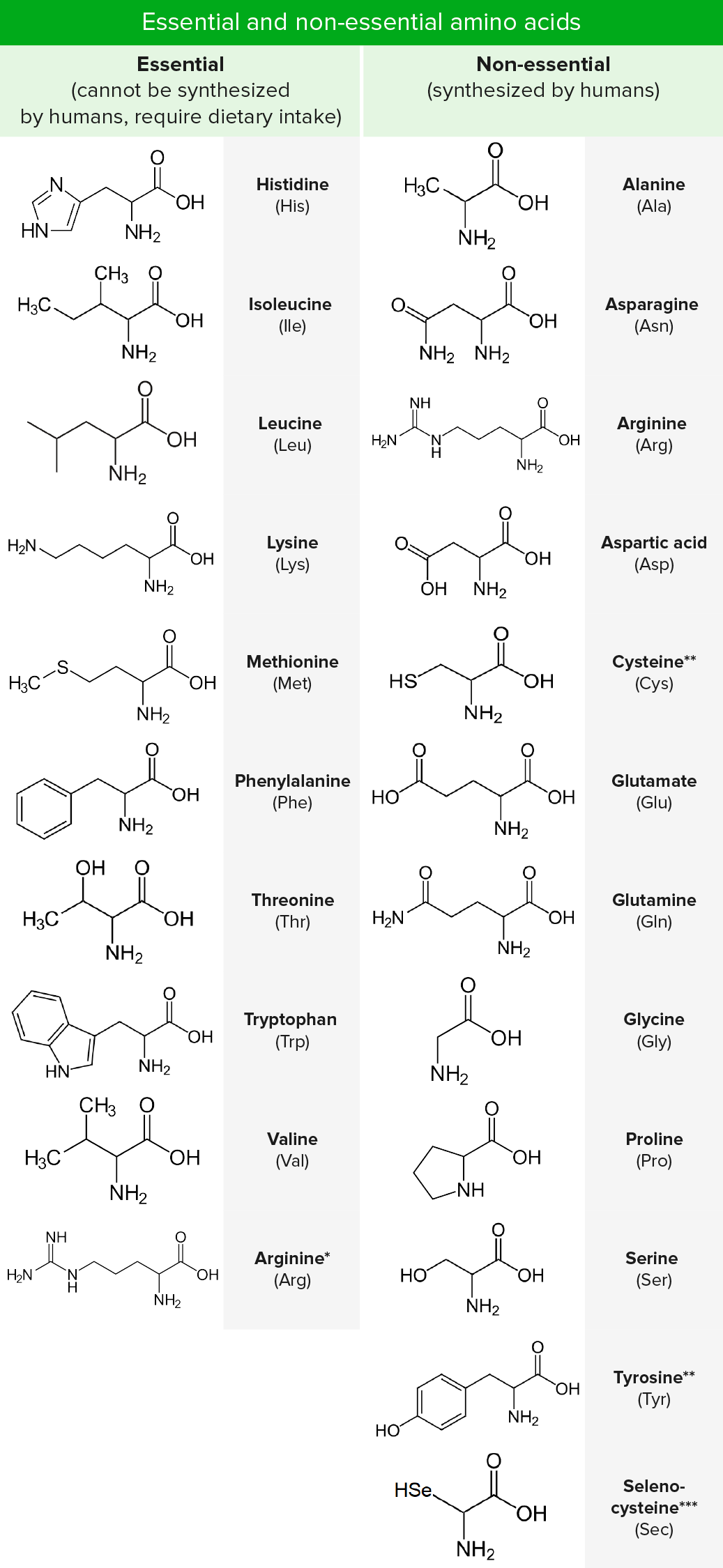
We examine this issue using data from a ribosomal footprinting assay in yeast. While there is great variation in ribosomal occupancy along even a single transcript, what determines a ribosome's occupancy is unclear. Both for understanding mechanisms of disease and for the design of transgenes, it is important to understand the determinants of ribosome velocity, as changes in the rate of translation are important for protein folding, error attenuation, and localization.
